Subream Breast MRI Lesion Lens

About
Summary
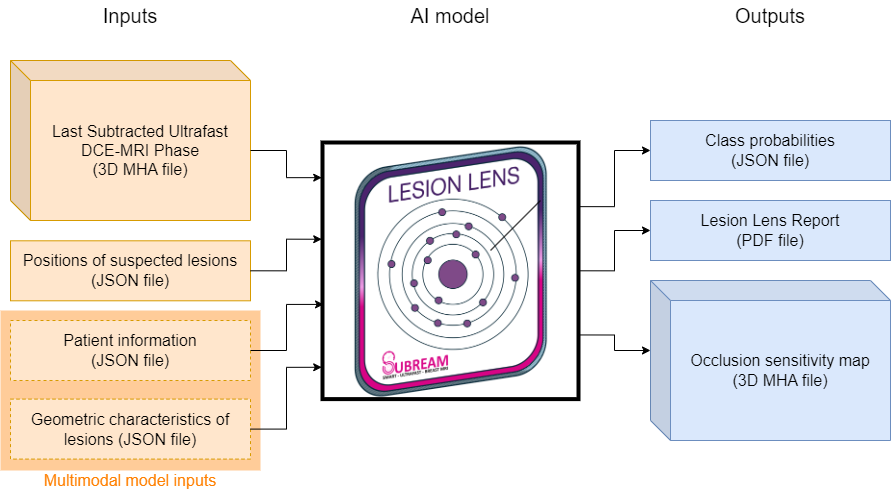
This Lesion Lens algorithm aims at classifying suspected breast lesions into malignant, benign or lymph nodes based on a subtracted image acquired with an ultrafast dynamic contrast-enhanced (DCE)-MRI sequence. Since suspected lesions are not automatically detected, their locations must be provided as an input to the algorithm.
This algorithm embeds two different deep learning models: a main multimodal model and a fallback image-only model.
The multimodal model considers three types of data modalities: imaging, categorical and scalar data. It uses transformer-like modules that rely on an attention mechanism in order to perform breast lesion classification. Three distinct attention modules are combined:
- A Vision Transformer (ViT) that projects images into a representation space shared with vector data.
- A “sieve” Transformer tasked to separate information that is shared across modalities from information that is exclusive to each modality.
- A final Transformer receives all exclusive and shared data to classify lesions.
The fallback image-only model is run when only the image and the location of the lesions are provided. This model is based on a standard DenseNet-121 and has been trained on the same cases as the multimodal model.
An automatic segmentation model based on a Swin-Unet is also embedded in LesionLens, allowing users to use the multimodal model without manually providing the geometric characteristics of the lesions (which represents most of the input scalar data).
This model was developed in 2022-2025 at the Geneva School of Health Sciences as part of the Smart and Ultrafast Breast MRI (SUBREAM) project funded by the Swiss Cancer Research (KFS-5460–08-2021-R) and the LesionLens project funded by the Fonds de Recherche et d’Impulsion (FRI) HES-SO (AGP: 133801). The project was approved by the Geneva Cantonal Ethics Committee (CCER) (Project-ID: 2019-00716). Informed consent was obtained from each patient for the re-use of anonymized breast imaging reports and MRI examinations.
Mechanism
Input¶
- Data type:
- [Required] Subtracted Image: Computed as the subtraction between the first and last phases of the ultrafast DCE-MRI sequence. File format is MHA or DICOM (in case of multiple DICOM files send them as a single ZIP archive).
- [Required] Lesion positions: 3D positions of the lesions in the image, and if available left and right nipple positions. File format is JSON. Left and right nipple positions are provided as "lesions" with names "nipple_left" and "nipple_right".
- [Optional] Patient data: Age of the patient and categorical fields (menopausal status, contraception, family risk, patient risk, BRCA and chemotherapy). Fill the required fields in the dialog menu or copy-paste the information in JSON format in the Code view.
- [Optional] Lesion geometry: Geometric characteristics of each lesion (bounding box center, bounding box size, volume, centroid, elongation, ellipsoid diameter and flatness). File format is JSON. Contains a
compute_lesion_geometryflag: When set to true, an automatic segmentation model embedded in LesionLens will compute these geometric characteristics automatically, replacing all existing geometric characteristics if present.
- Target population: Patients who underwent breast MRI examination with a protocol including Ultrafast MRI sequence with enhanced breast lesions.
Please refer to interface information and warning sections for details on JSON files content.
Output¶
- Class probabilities: probability for each class (benign, lymph node or malignant)
- Interpretability: occlusion maps are computed based on the 3D image and the model predictions.
- PDF report: A report summarizing the most relevant information for the cases and the suspected lesions.
Model type: Transformer-based neural network.
Training data location and time-period: Training data were collected retrospectively between 2019 and 2023 at a clinical center in Geneva (Switzerland) acquired with 3T Philips Ingenia. The dataset comprised 987 lesions (280 benign, 121 malignant lesions, and 586 benign lymph nodes) identified from 301 breast MRI examinations (patient mean age 54.8 ± 12.7).
External validation: External validation was made with data collected at clinical center in Neuchâtel (Switzerland) acquired with 3T Siemens Vida. The external dataset comprised 248 lesions (45 benign, 107 malignant lesions, and 96 benign lymph nodes) identified from 117 breast MRI examinations (patient mean age 52.7 ± 15.4).
Publicly available results¶
Due to ethical restrictions, results applied on data of the paper cannot be made public. As a result, we exemplify the algorithm on public data. Note that these datasets do not use ultrafast MRI sequences but rather DCE MRI alike sequences. As a result, while provided results highlight the algorithm success, it may fail in other situations, especially if lesion and patient characteristics are not provided (i.e. reliance only on image):
- Result 1 (GC id: d8bcce62-136e-4122-a6d8-fb186e6c0b83): Case #AMBL-003 from the Advanced-MRI-Breast-Lesions datasaet (Daniels et al. 2024). This case showcases a malignant lesion (invasive ductal carcinoma) in the right breast. Lesion characteristics were computed based on the provided segmentation in the AMBL dataset.
- Result 2 (GC id: eb5c7be2-087e-4a3d-99a2-4b5b2fc6c5fe): Case #294 from FastMRI Breast MRI dataset (Solomon et al. 2025). Another case of invasive ductal carcinoma. We also included an example of two lymph nodes on left side to illustrate the AI capability. Lesion segmentation was performed since the FastMRI dataset does not include them.
Daniels, D., Last, D., Cohen, K., Mardor, Y., & Sklair-Levy, M. (2024). Standard and Delayed Contrast-Enhanced MRI of Malignant and Benign Breast Lesions with Histological and Clinical Supporting Data (Advanced-MRI-Breast-Lesions) (Version 2) [Data set]. The Cancer Imaging Archive. https://doi.org/10.7937/C7X1-YN57
Solomon, E., Johnson, P. M., Tan, Z., Tibrewala, R., Lui, Y. W., Knoll, F., ... & Heacock, L. (2025). fastMRI Breast: A publicly available radial k-space dataset of breast dynamic contrast-enhanced MRI. Radiology: Artificial Intelligence, 7(1), e240345. https://doi.org/10.1148/ryai.240345
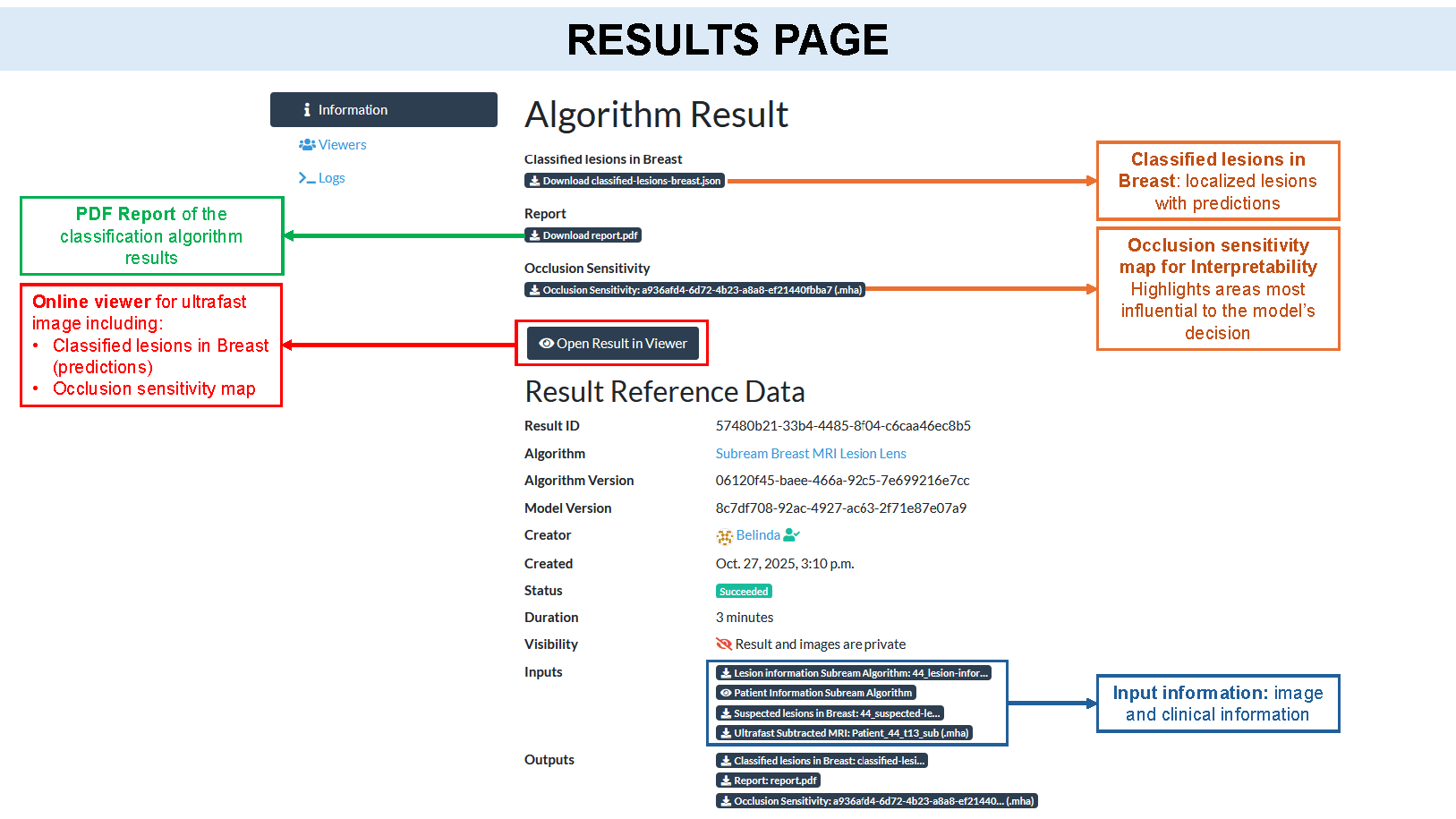
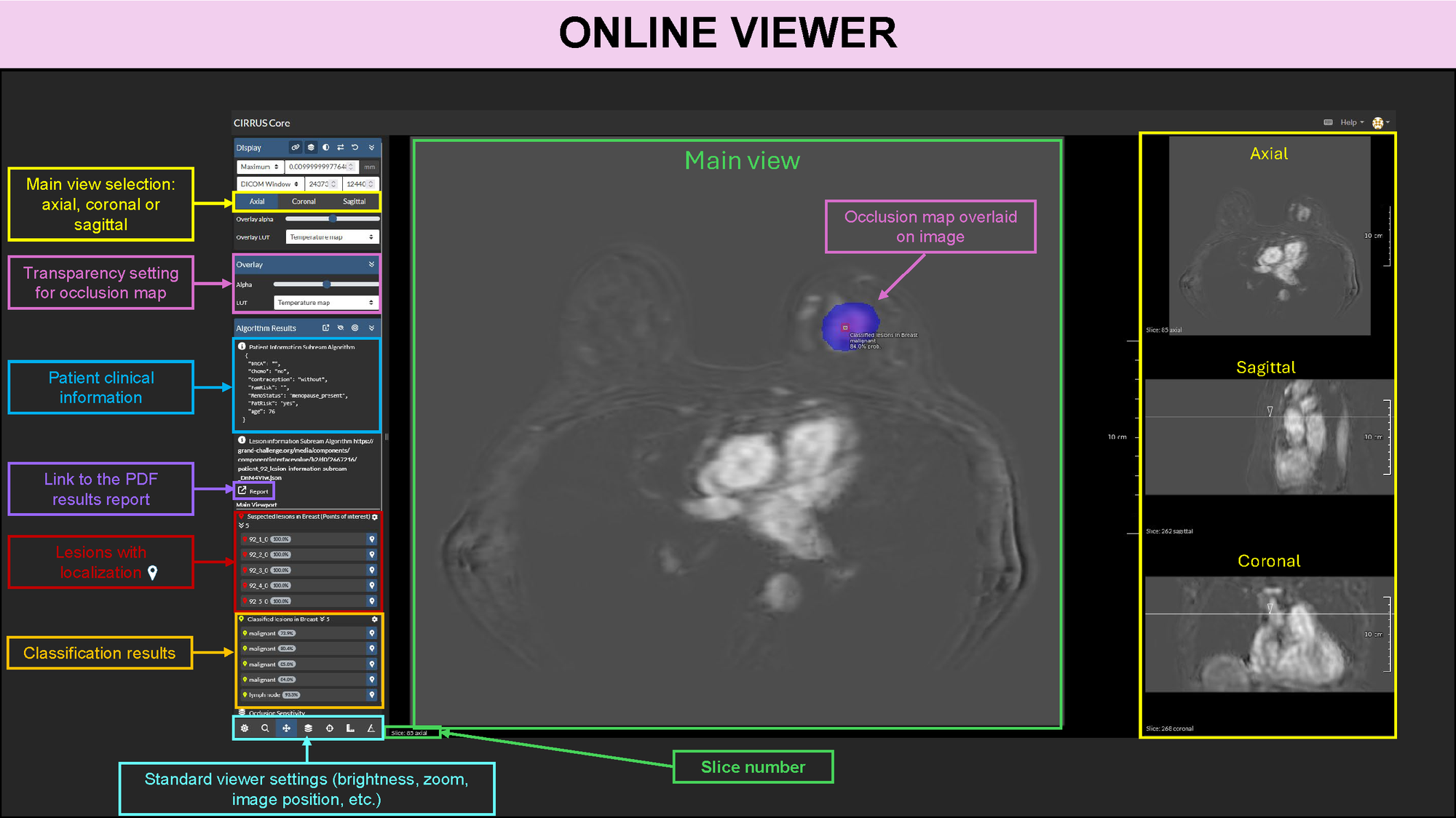
Interfaces
This algorithm implements all of the following input-output combinations:
| Inputs | Outputs | |
|---|---|---|
| 1 | ||
| 2 |
Validation and Performance
Multimodal model
| AUROC (%) | Sensitivity at EER (%) | Specificity at EER (%) | Specificity at 90% Sensitivity | |
|---|---|---|---|---|
| Local | 92.8 (± 2.7) | 86.7 (± 2.8) | 86.3 (± 3.0) | 70.8 (± 14.4) |
| External | 78.9 | 70.9 | 71.6 | 38.3 |
Unimodal model
| AUROC (%) | Sensitivity at EER (%) | Specificity at EER (%) | Specificity at 90% Sensitivity | |
|---|---|---|---|---|
| Local | 86.3 (± 2.5) | 76.1 (± 6.6) | 78.3 (± 4.1) | 53.1 (± 14.6) |
| External | 73.8 | 71.8 | 72.3 | 25.5 |
EER: Equal Error Rate
Local: Results for the local dataset were obtained using 5-fold cross-validation, and are reported as mean (+/- standard deviation). https://doi.org/10.1016/j.compbiomed.2025.109721
External: External validation with clinical center at Neuchâtel (Switzerland)
Uses and Directions
For research use only. This algorithm is a prototype and was developed for research purposes only. It is therefore not intended for direct clinical application.
- Expected benefits: The integration of ultrafast DCE-MRI with clinical data may improve the classification of breast lesions, potentially supporting earlier and more accurate decision-making.
- Target population and use case: Patients undergoing breast MRI for any clinical indication. The model is designed to support radiologists in the assessment of breast lesions detected on MRI.
- General use: This model is intended to assist radiologists or, under supervision, MRI radiographers, in the classification of breast lesions using ultrafast DCE-MRI combined with clinical parameters. It is not a diagnostic tool and should not independently guide patient management. The model is designed to complement, not replace, the radiologist’s interpretation and clinical judgment.
- Appropriate decision support: The model analyzes ultrafast DCE-MRI and relevant clinical information to provide a classification of breast lesions. The radiologist reviews these results in the context of the patient’s full clinical profile and determines the most appropriate diagnostic and follow-up recommendations.
- Before using this model: The algorithm should be validated retrospectively and prospectively on diagnostic cohorts that accurately reflect the target population to confirm performance and generalizability in the intended clinical setting.
- Safety and efficacy evaluation: To be determined.
Warnings
- Risks: Even if used appropriately, the model may produce incorrect predictions. Outputs should always be interpreted with caution. The position of the lesion in the PDF report may not be reliable, due to the use of an illustrative image and the position of the breast in the MR image. However the distance to the nipple is accurately computed based on the nipple and lesion positions.
- Inappropriate Settings: This model was trained exclusively on Ultrafast MRI sequences. It has not been trained or validated on standard DCE-MRI protocols and may perform inaccurately outside its training domain.
- Clinical Rationale: The model provides visual evidence for the predicted lesions based on occlusion maps. While these maps support model interpretability, they are not always reliable as they may not capture all the factors influencing the model’s predictions. Clinical end users are expected to place the model output in context with other clinical information to make final determination of diagnosis.
- Inappropriate decision support: This model may not be accurate outside of the target population.
- Generalizability: This model was primarily developed with ultrafast DCE-MRI data from clinical center in Geneva and tested with data from clinical center in Neuchatel (Switzerland). Do not deploy this model in other institutions or populations without prior evaluation.
- Discontinue use if: Conditions for stopping model use are still under evaluation and will be defined in future versions.
Common Error Messages
Errors
| Error message | Solution |
|---|---|
| Your image have N dimensions but expected a 3D image | Make sure your image has 3 dimensions (width, height, depth) |
Your Suspected lesions in Breast file does not contain any lesion. |
Make sure your Suspected lesions in Breast follows the expected pattern and contains the positions of the lesions |
| Incorrect lesion data configuration [...] | Depending on the rest of the error message, you may have lesions in your Lesion information that are either missing or incorrectly named from the Suspected lesions in Breast file (or vice-versa). In all cases, make sure the list of lesion IDs in both files are the same |
| Suspected lesions in Breast. JSON does not fulfill schema [...] | Make sure the JSON file for suspected lesions and their positions follows the required schema. For example, one of your positions may have an incorrect number of coordinates (exactly 3 coordinates per position are required) |
Lesion ... located at [...] is out of bounds. Coordinate(s) out of bounds: |
Make sure the position of the lesion falls within the image bounds. |
| Unsupported number of nipples found: 1 | When possible, provide the position for both nipples. Make sure they are correctly named. Supported names start with "nipple" and end with either "left"or "right". Names are not case sensitive. If only one nipple position is available, remove it from the list of lesion positions, as the report won't be able to handle this case. |
| Left and right nipples have the same X position (left/right axis) | Check and update the position of nipples. |

