August 2023 Cycle Report
Published 10 Aug. 2023
Experimental client-side viewer🧪¶
We're excited to introduce a new experimental feature on grand-challenge.org: the Client Rendered View Item for Pathology. This feature utilizes OpenSeaDragon to render pathology images on the client side, enhancing responsiveness compared to the traditional server-rendered cirrus view item. While in its experimental phase, this feature currently supports zoom and pan tools, allowing users to analyze images more smoothly. Additionally, it operates as a linked view item, ensuring synchronization between (client- and server-rendered) view items. The view item does not yet support advanced features like annotations or overlays.
To enable this feature a user needs to create a custom hanging protocol with a viewport object containing the property: "specialized_view": "clientside". This will create a viewport using the experimental feature. Then, select the custom hanging protocol in a viewer configuration to start using it.
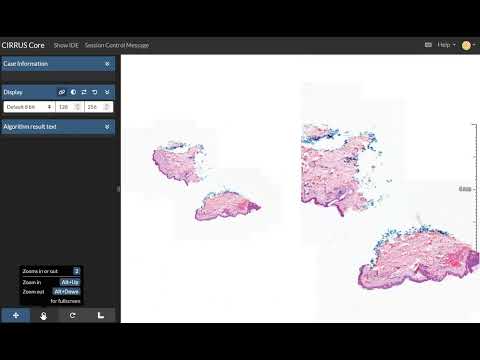
See the video below for a demo of the client-side view item on the left versus a server-side rendered view item on the right.
Combined Leaderboards 🏆¶
This cycle sees the introduction of a new type of Challenge leaderboard: the combined leaderboard. This new leaderboard type allows observing a user's algorithm(s) performance across multiple phases or tasks. Combined leaderboards rank performance via a combination of the ranks of several challenge phases.

Challenge admins can add the new leaderboards via the new button under the section 'Combined Leaderboards' in the admin interface. The current options for the combination method are the mean, median or sum of the ranks.
Improved GitHub Repository Integration  ¶
¶
The GitHub Repository Integration has been improved with a new integration workflow. Searching for installed repositories has also been added so the form now works for people who are admins for more than 100 repositories in their organisation. You can now find all the settings related to GitHub integration in a single form:

Overlapping segmentations¶
CIRRUS and grand-challenge now support a new type of multi-class overlays which can be used to show multi-category masks on top of an image, where regions of different categories are allowed to overlap.
If you want to use this new feature, you must add an extra dimension to the mask you create. For example, for a 3D mask, you would generate a 4D image where the first dimension will represent the mask value. The resulting 4D mask must be written as a stacked binary image. So the mask, representing "category 2" would be referred to by the (numpy) index mask[2, :, :, :]. The mask values themselves must be of type uint8 and be either 0 or 1 (binary).
CIRRUS can render such a mask as a multicategory mask with overlapping regions. You can make these regions more visible by playing with the overlay-alpha or disabling different segments in the OverlayPlugin.
Provide feedback regarding our Image Viewer ⌨️¶
The RSE team is always looking for ways to improve our software, and we are wondering what the biggest shortcomings are of the Cirrus viewer (the application you see when viewing images, for example in a reader study or when viewing algorithm results). This is your chance to provide feedback and tell your grievances and improvement tips! Fill in the questionnaire here, it will only take a few minutes, and will be much appreciated!
Cover photo by Viswa Teja on Unsplash