Spring Newsletter 2026
Published 2 March 2026
🛠️ New features of Grand Challenge
Welcome to the Early Spring Edition of Grand Challenge Insights!
In this blog, we will share the latest updates and highlights from our Grand Challenge community.

Viewer ecosystem updates
When opening images, the viewer has to go through several checks and setup steps before anything is shown, which can cause noticeable waiting time. In this cycle, we simplified the ecosystem of the viewer by removing user-specific setup and older access requirements. These changers prepares the viewer ecosystem for future changes that should greatly improve start-up timing.
Automatic hanging protocol selection
You can now automatically select suitable hanging protocols per set when having a varying number of images and other data, enabling you to vary your input data per case. Hanging protocols define how images and other data are arranged in the viewer. This automatic selection is applied under specific conditions when the number of items to be shown differs between cases. The exact conditions for when a hanging protocol is selected automatically are described in the documentation.
 Generated with DALL-E.
Generated with DALL-E.
DICOM upload and viewing fixes
We started on refactoring the image upload functionality to better support DICOM and improve maintainability of the existing code. In addition, several DICOM-related issues were fixed, including problems with uploading DICOMs with multiple files and scaling issues. Additionally, small example snippets for reading and writing DICOM were also added to algorithm and challenge starter kit templates.💡 Blog posts¶
"December 2025 Cycle Report"¶
Read all about the platform improvements from our RSE team in December, including work on updates to DICOM upload and viewing and faster viewer startup.
"February 2026 Cycle Report"¶
Read all about the platform improvements from our RSE team in January and February, including automatic hanging protocol selection and preparation for faster viewer launches.
"March 2026 Cycle Report"¶
Read all about the platform improvements from our RSE team in January and February, including automatic hanging protocol selection and preparation for faster viewer launches.
🖊️ Use case: Running multi-reader annotation studies on Grand Challenge¶
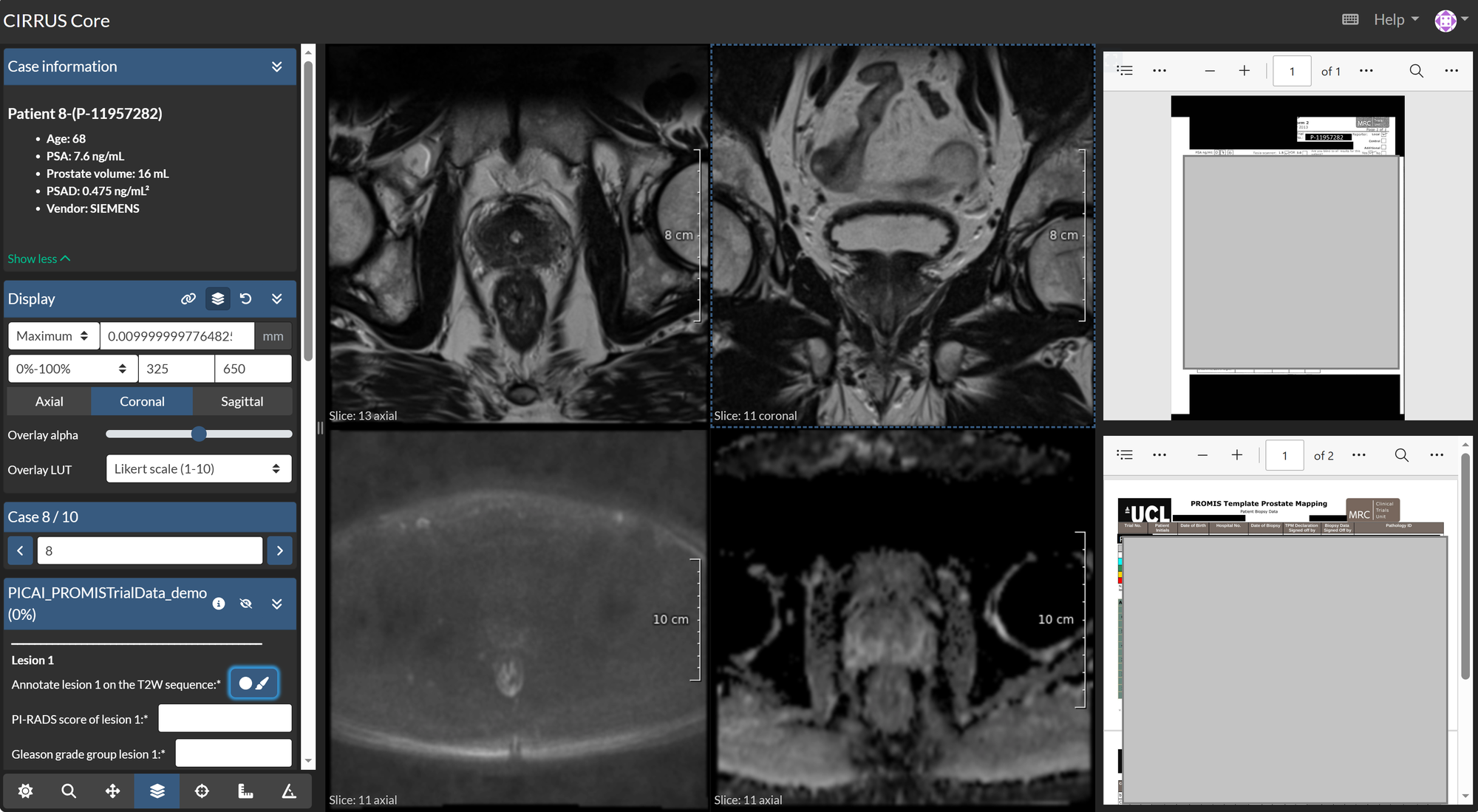
For a follow-up study on PI-CAI, annotations were required. For this project, multiple readers were invited, each reviewing three MRI modalities (T2W, ADC, HBV) with the pathology and radiology reports for each patient case. Using Grand Challenge, annotators could access imaging data and reports within a single environment, perform lesion annotations, and record the corresponding pathology and radiology scores in a structured manner. Such features can be used also in other medical domains such as pathology when reviewing pathological slide images, or images from other modalities (CT, PET, etc.). Both the reports and the imaging data can be minimized and maximized to occupy the full screen.
Because the readers were distributed across sites, local reading was not feasible. Additionally, VPN-based access to the hospital network was not possible for devices not managed by the hospital. Therefore, setting this annotation process in the Grand Challenge platform made it possible to run the study centrally, without requiring local software installation or network access to the hospital storage systems.
Using this structure, patient data has been annotated by readers, with enforced answer formats and no per-reader software configuration. After completion, the annotations can be downloaded for local storage or retained on Grand Challenge for use in downstream algorithms.
Highlighted Challenges¶
🥅 Goal: The BEETLE benchmark focuses on semantic segmentation of breast cancer histopathology in H&E whole-slide images. It provides a multicentre, multiscanner dataset with pixel-level annotations, aiming to drive progress toward robust, generalizable segmentation models that can support automated biomarker quantification in breast cancer.
Task: Multiclass semantic segmentation in breast cancer histopathology slides
Preprint: https://arxiv.org/abs/2510.02037
🔦 Publication highlight¶
We congratulate the DRAGON benchmark organisers with their publication entitled "Leveraging open-source large language models for clinical information extraction in resource-constrained settings". This study tested 9 open-source artificial intelligence (AI) language models on extracting information from Dutch medical records on 28 clinical tasks. The authors concluded that "modern generative AI can support clinical data analysis in Dutch with minimal setup, offering a flexible alternative to traditional models that require large amounts of labeled training data"
Builtjes, L., Bosma, J., Prokop, M., van Ginneken, B., & Hering, A. (2025). Leveraging open-source large language models for clinical information extraction in resource-constrained settings. JAMIA open, 8(5), ooaf109.
https://doi.org/10.1093/jamiaopen/ooaf109
🔦 Highlighted algorithms¶
Deep Learning Image Quality for Prostate MRI
This algorithm is a deep learning classifier designed to automatically assess the quality of axial T2-weighted prostate MRI scans. The model was trained to distinguish between low- and high-quality images and to provide a continuous score reflecting overall image quality. To enhance interpretability, the algorithm also generates class-specific heatmaps that visualize which regions influenced the prediction.
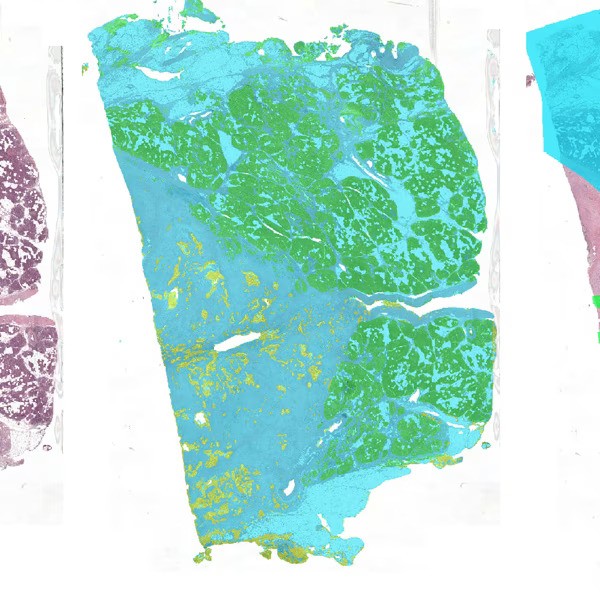
PDAC Tumor Segmentation
This is a segmentation algorithm for Pancreatic Ductal Adenocarcinoma in pancreatic whole-slide images. It applies a three step pipeline for segmentation and outputs:- a mask with the tissue, healthy epithelium, and tumor epithelium
- a mask with the tumor bulk area

